Resen
Resen, the REproducable Software ENvironment, is a tool that enables scientific data analysis and reproducibility by:
- providing pre-installed community developed software and tools (no more hours wasted on installing software), and
- packaging data analysis code and results in an isolated platform independent software environment that can be shared and published to cite-able online repositories (such as Zenodo).
The goal of resen is to lower the barrier to entry for performing data analysis and facilitate reproducibility (resen is one necessary, but not sufficient tool). Using resen, scientists should allow you to spend less time installing packages and fixing hard-coded file paths so you can get to your scientific interpretation and results faster.
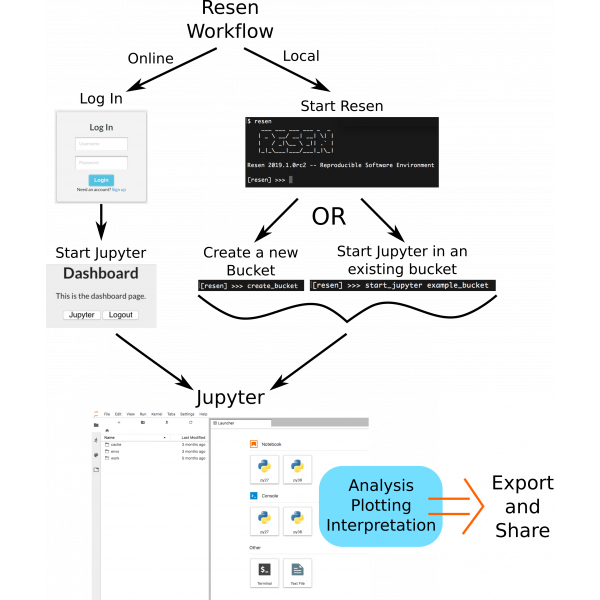
Using Resen
Resen can be used both online through the InGeO server and locally on your own computer. The option you choose should depend on how familiar you are working in python in a jupyter environment and the amount of data/computational resources you will require. Both options give you access to the same reproducable environment.
Try it Online - Make an account and explore resen online. Python novices and people less familiar with resen should start here. Tutorials are available to help you get started and you can even upload your own (small) datasets to start producing results immediately.
Install Resen and Run Locally - Install resen run it locally on your own computer. This is a good option if you would like to take advantage of local computational resources or would like to use large amounts of data.
Key concepts
When working with resen, you will work within a "bucket". A bucket is an analysis environment that based on a core and any data and software that is mounted in. A core is an image with pre-installed software and tools. Once a bucket is created, access to the bucket is provided via JupyterLab in the web browser on your computer. If a scientists is working on 3 papers at a time, they can have a bucket for each, each with a different configuration and installed software. When analysis is complete or ready to be published, the bucket can be exported from resen and uploaded to a cite-able repository.
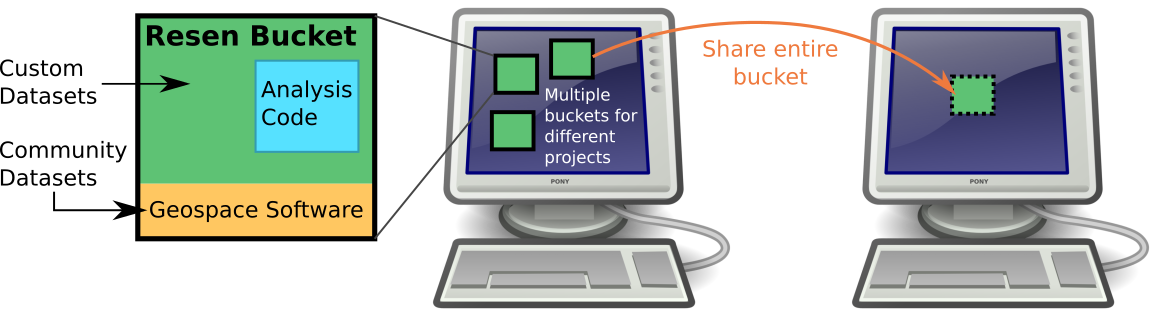
How it works
Resen leverages existing robust container technology. Containers are portable, use less system resources than virtual machines, and provide an isolated software environment within which code can be executed. Portability means that containers can be transferred to a completely different system and it will behave identically. For example, a container can be run on linux, macos, or windows! In addition, the isolated software environment aspect enables reproducibility.
For resen, we have chosen to use docker containers because of its wide commercial adoption and open source codebase.
Features
- Tutorials are provided to facilitate learning about community tools and working with them.
- Resen is open source, meaning you can contribute to resen or build your own custom cores
- Via community tools, resen includes APIs for easily accessing some common geoscience data sets.
- You can import and use your own data with Resen.
- Resen can either be run locally or in a server environment in the cloud
The punchline
By using containers, scientists can seamlessly share analysis code and results with each other without the frustration of trying to set up identical software environments and changing file paths. With containers, scientists can trivially import and reproduce previously published analysis results!
To learn more about resen and report bugs or issues, visit us on GitHub.